New models for studying SARS CoV-2 Spike/human ACE2 interaction
Severe acute respiratory syndrome coronavirus 2 (SARS-CoV-2) is the etiologic agent responsible for the recent coronavirus disease 2019 (COVID-19) pandemic. Productive SARS-CoV-2 infection relies on viral entry into cells expressing angiotensin-converting enzyme 2 (ACE2). Indeed, viral entry into cells is mostly mediated by the early interaction between the viral spike protein S and its ACE2 receptor. The S/ACE2 complex is, thus, the first contact point between the incoming virus and its cellular target; consequently, it has been considered an attractive therapeutic target.
To further characterize this interaction and the cellular processes engaged in the entry step of the virus, V. Parissi (MobileVIR team, Bordeaux) developed a strong collaboration network between MFP lab, Vect’UB (V. Guyonnet-Dupeyrat), IMPACT platform in Nantes (C. Charlier and P. Weigel), IINS (A. Favereaux) and BioSIM, University of Porto. This collaborative work led to set up various in silico, in vitro and in cellulo approaches that allow to specifically monitor the S/ACE2 association.
- In silico monitoring of the SARS-CoV-2 S-RBD/ACE2 interaction.
A computational model of the SARS-CoV-2 S/ACE2 complex was developed to provide data about the dynamics of the interaction. The simulation allowed to precisely model the structural interfaces within the complex and isolate the Receptor Binding domain of S (RBD) (Fig1A). In addition, the analysis of the interfacial hydrogen bounds was generated by molecular dynamics (Fig1B).

- In vitro monitoring of the SARS-CoV-2 S-RBD/ACE2 interaction.
In this work, IMPACT platform contributed to the S/ACE2 complex characterization using biochemical and biophysical approaches such as AlphaLISA (Fig2A) and biolayer interferometry (BLI) (Fig2B) binding assays. This led to determine the kinetic parameters of the S/ACE2 association and dissociation steps.
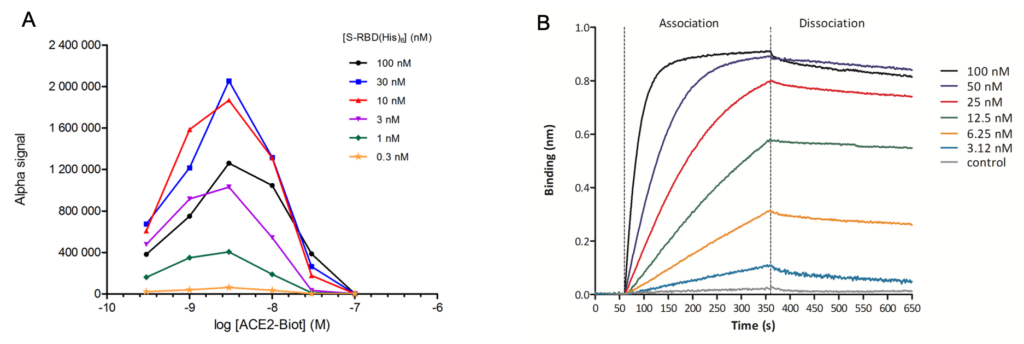
Fig 2. In vitro monitoring of the SARS-CoV-2 S-RBD/ACE2 interaction. SARS-CoV-2 S-RBD(His)6 and human ACE2-Biot interaction have been first monitored using AlphaLISA assay. Cross titration experiments of ACE2-Biot (0.3 to 100 nM) against S-RBD(His)6 (0.3 to 100 nM) using anti-6X His acceptor beads and streptavidin donor acceptor beads are reported in (A). The dynamics of the S-RBD/ACE2 interaction was monitoring using Bio-Layer Interferometry. Streptavidin biosensors were loaded with 1 µM of ACE2-Biot. After the baseline step (60 s, each loaded biosensor was dipped into serial dilutions of S-RBD(His)6 (from 3.12 nM to 100 nM) for 300 s for the association step. The dissociation step of the bound S-RBD(His)6 was monitored by dipping the biosensor back into the reaction buffer for 300 s (B). Kinetic binding parameters (kon, koff) and the equilibrium dissociation constant (KD) were determined.
- In cellulo monitoring of S/ACE2 mediated viral entry and cell fusion
In parallel to these in vitro approaches, a novel computational model of the SARS-CoV-2 S/ACE2 complex was developed to provide data about the dynamics of the interaction. For the in cellulo part of the project, transduction assays were performed using SARS-CoV-2 pseudotyped lentiviral vectors and HEK293T-ACE2 cell lines generated in-house. This allowed to recapitulate the early replication stage of the infection mediated by the S/ACE2 interaction and to detect cell fusion induced by the interaction. Finally, a cell imaging system was set up to directly monitor the S/ACE2 interaction in a cellular context, and a flow cytometry assay was developed to quantify this association at the cell surface.

- Together, these different approaches are available for both basic and clinical research aiming to characterize the entry step of the original SARS-CoV-2 strain and its variants as well as to investigate the possible chemical modulation of this interaction.
- All these models will help in identifying new antiviral agents and new chemical tools for dissecting the virus entry step as currently ongoing in the team.
Reference:
2021 – In Silico, In Vitro and In Cellulo Models for Monitoring SARS-CoV-2 Spike/Human ACE2 Complex, Viral Entry and Cell Fusion. Lapaillerie D., Charlier C., Fernandes H.S., Sousa S.F., Lesbats P., Weigel P., Favereaux A., Guyonnet-Duperat V. and Parissi V. Viruses. 13(3):365. doi.org/10.3390/v13030365.
